DREAMS: deep read-level error model for sequencing data applied to low-frequency variant calling and circulating tumor DNA detection, Genome Biology
Por um escritor misterioso
Last updated 23 maio 2024
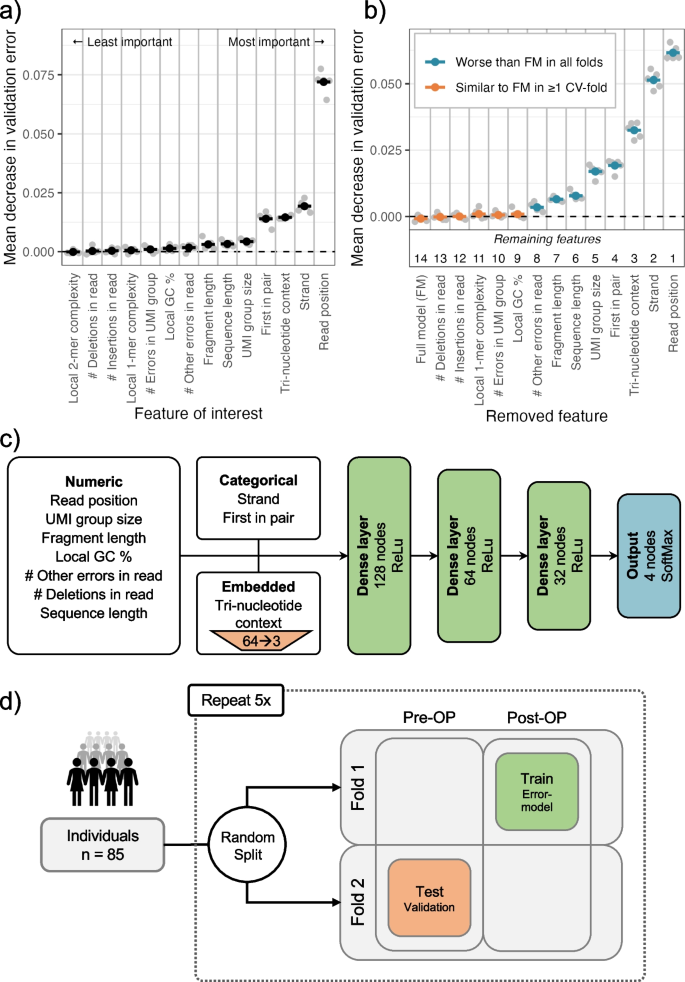
Circulating tumor DNA detection using next-generation sequencing (NGS) data of plasma DNA is promising for cancer identification and characterization. However, the tumor signal in the blood is often low and difficult to distinguish from errors. We present DREAMS (Deep Read-level Modelling of Sequencing-errors) for estimating error rates of individual read positions. Using DREAMS, we develop statistical methods for variant calling (DREAMS-vc) and cancer detection (DREAMS-cc). For evaluation, we generate deep targeted NGS data of matching tumor and plasma DNA from 85 colorectal cancer patients. The DREAMS approach performs better than state-of-the-art methods for variant calling and cancer detection.

DREAMS: Deep Read-level Error Model for Sequencing data applied to

General illustration of our approach. (a) Distribution of observed

Analytical validation of an error-corrected ultra-sensitive ctDNA
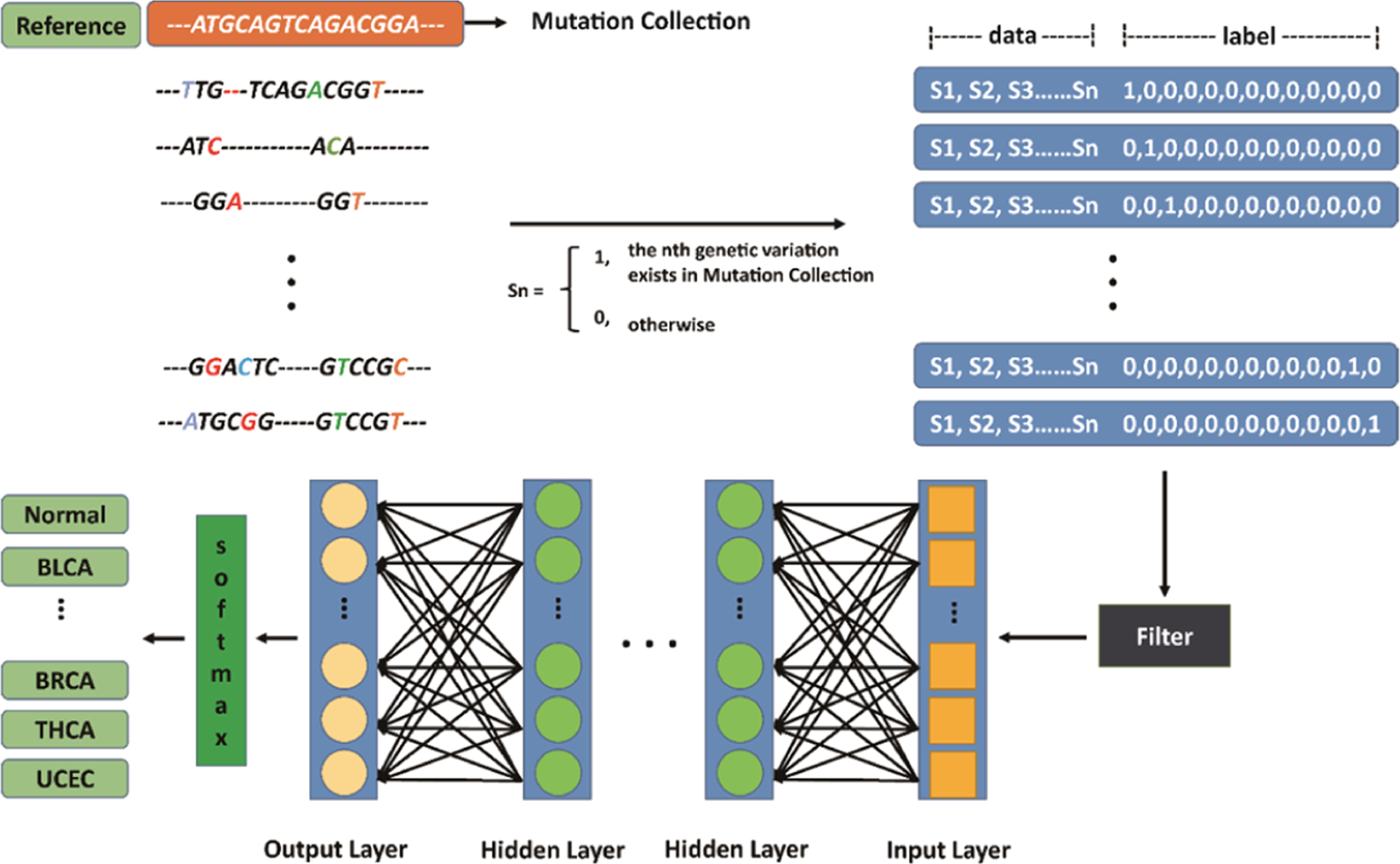
Identification of 12 cancer types through genome deep learning
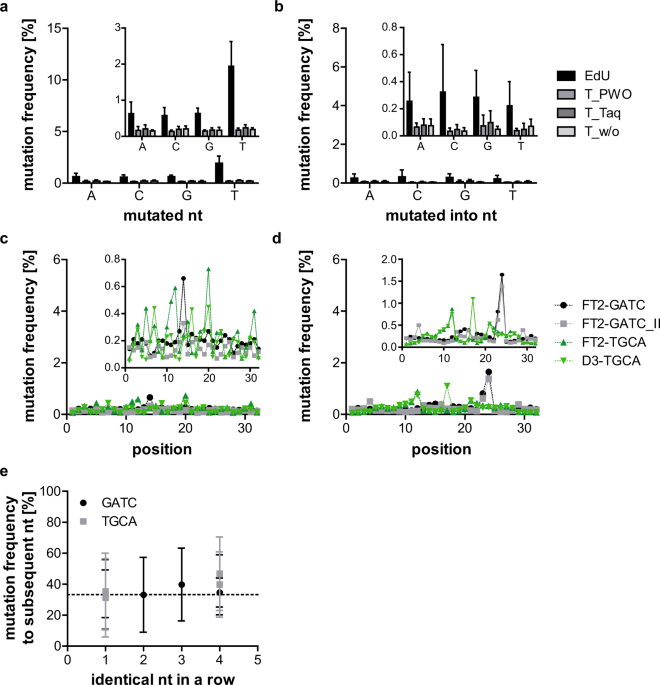
Systematic evaluation of error rates and causes in short samples
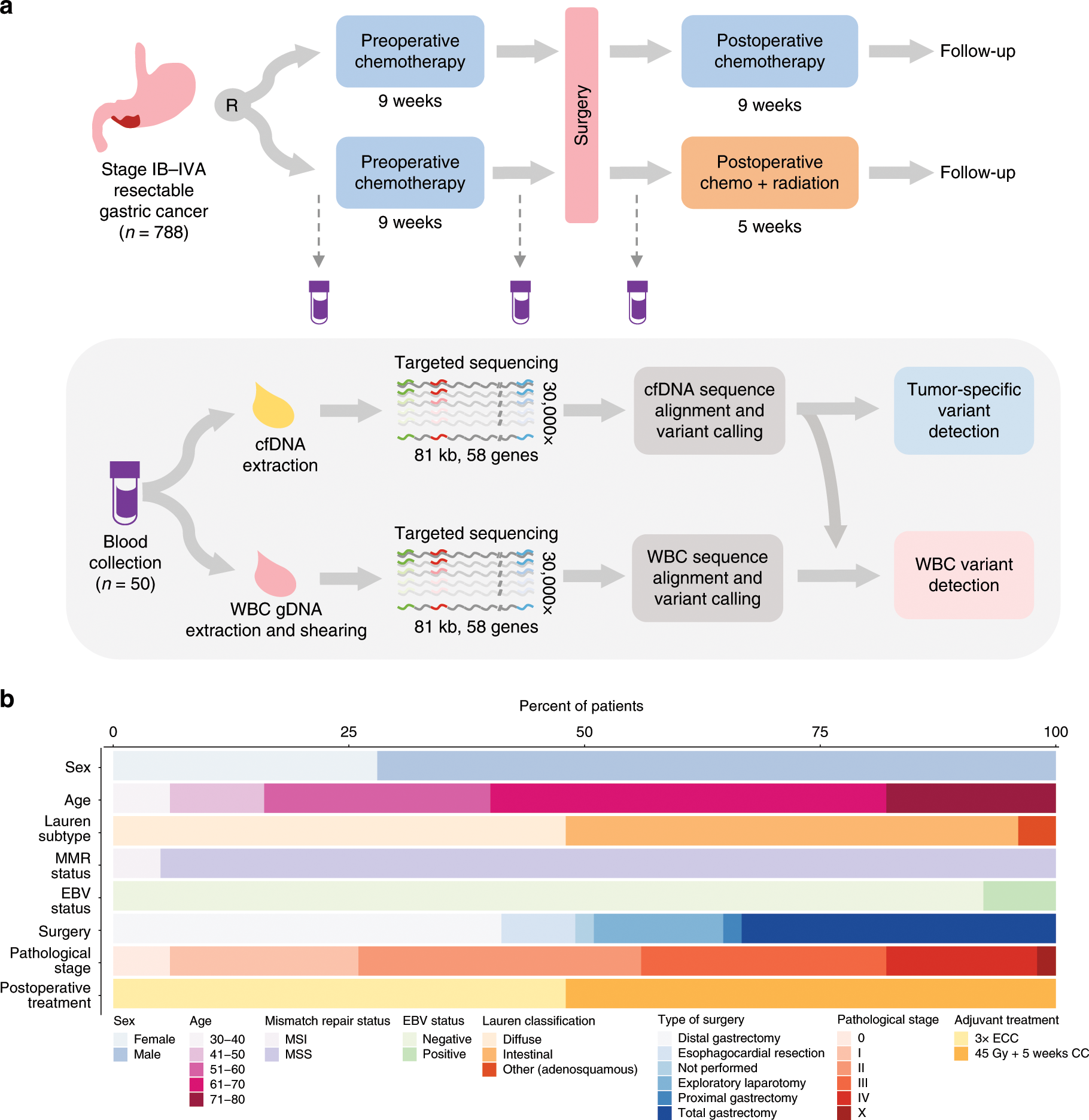
White blood cell and cell-free DNA analyses for detection of

PDF) DREAMS: deep read-level error model for sequencing data

The changing face of circulating tumor DNA (ctDNA) profiling
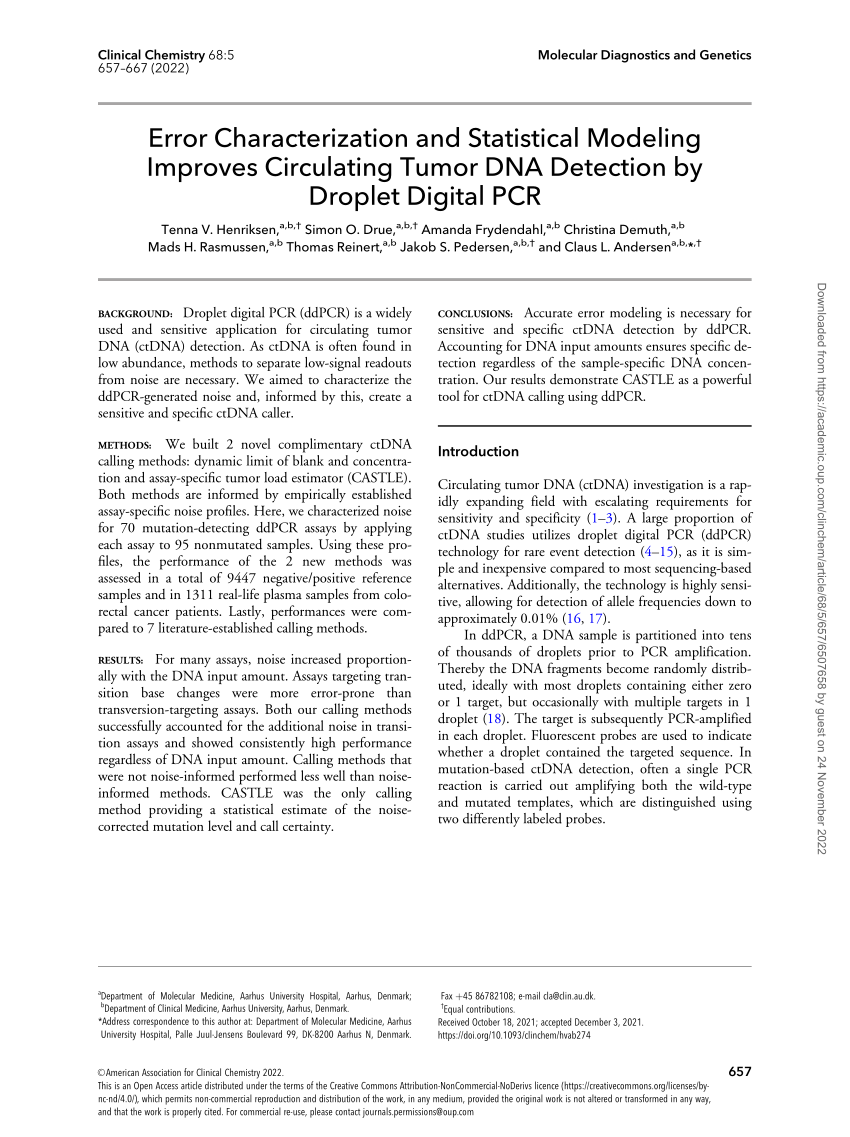

PDF) Error Characterization and Statistical Modeling Improves

Machine learning guided signal enrichment for ultrasensitive
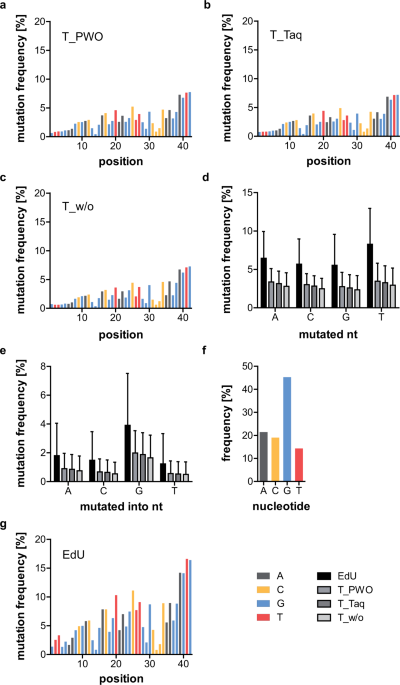
Systematic evaluation of error rates and causes in short samples

Optimizing Cancer Genome Sequencing and Analysis - ScienceDirect
Recomendado para você
-
 Picture memes vksFDjFX6 by I_eat_ass____2017 - iFunny Brazil23 maio 2024
Picture memes vksFDjFX6 by I_eat_ass____2017 - iFunny Brazil23 maio 2024 -
 OC snubbed from an /x/ thread, SCP Foundation23 maio 2024
OC snubbed from an /x/ thread, SCP Foundation23 maio 2024 -
 Why my pp hard : r/DankMemesFromSite1923 maio 2024
Why my pp hard : r/DankMemesFromSite1923 maio 2024 -
 scp 7143 j23 maio 2024
scp 7143 j23 maio 2024 -
 Emolog by Kyungmoo Min23 maio 2024
Emolog by Kyungmoo Min23 maio 2024 -
 SCP 6034 - The War With Another Dimension23 maio 2024
SCP 6034 - The War With Another Dimension23 maio 2024 -
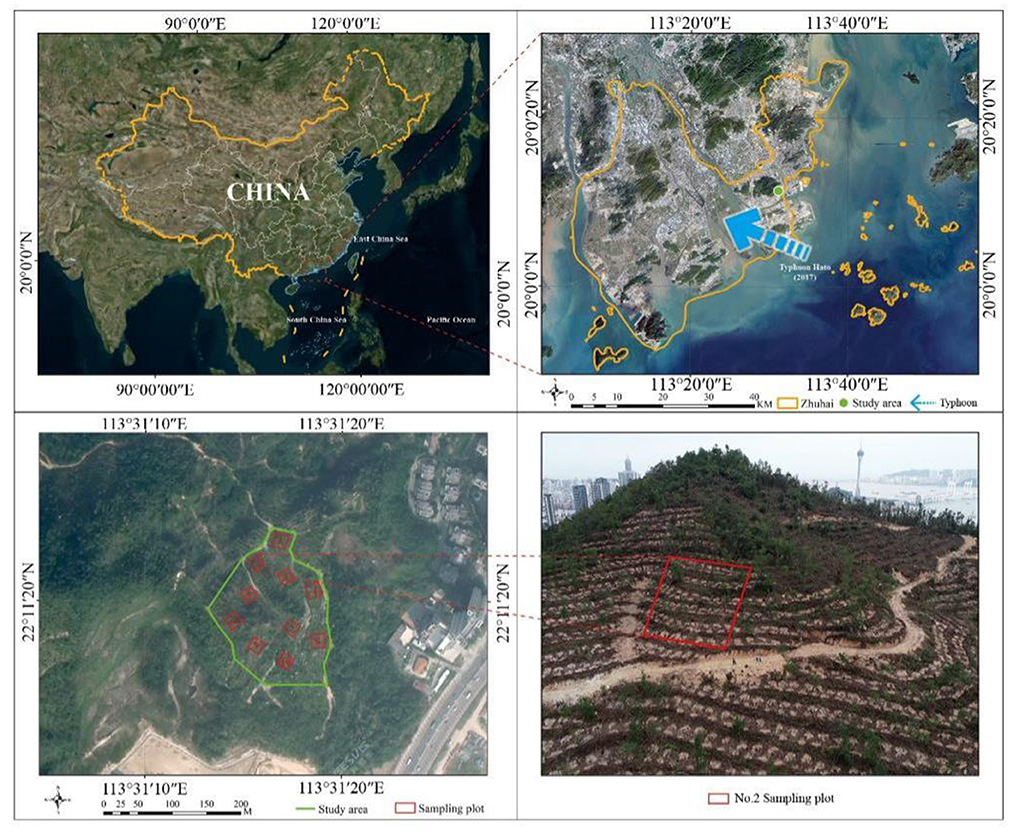 Frontiers Growth patterns and environmental adaptions of the tree species planted for ecological remediation in typhoon-disturbed areas—A case study in Zhuhai, China23 maio 2024
Frontiers Growth patterns and environmental adaptions of the tree species planted for ecological remediation in typhoon-disturbed areas—A case study in Zhuhai, China23 maio 2024 -
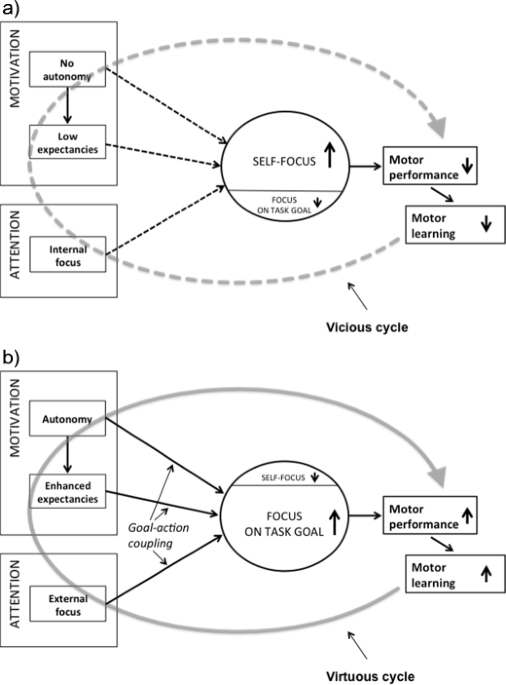 Optimizing performance through intrinsic motivation and attention for learning: The OPTIMAL theory of motor learning23 maio 2024
Optimizing performance through intrinsic motivation and attention for learning: The OPTIMAL theory of motor learning23 maio 2024 -
 Closing door close up hi-res stock photography and images - Alamy23 maio 2024
Closing door close up hi-res stock photography and images - Alamy23 maio 2024 -
Creature's Teeth Roblox Item - Rolimon's23 maio 2024
você pode gostar
-
Chainsaw Man (English Dub) MISSION START - Watch on Crunchyroll23 maio 2024
-
 Eric Clapton Tears In Heaven Sing Along Lyrics23 maio 2024
Eric Clapton Tears In Heaven Sing Along Lyrics23 maio 2024 -
antiquariorecifeoficial23 maio 2024
-
 Assassin's Creed III Remastered Switch update out now (version 1.0.2)23 maio 2024
Assassin's Creed III Remastered Switch update out now (version 1.0.2)23 maio 2024 -
 PlayStation Plus – PlayStation.Blog23 maio 2024
PlayStation Plus – PlayStation.Blog23 maio 2024 -
 Aquele Jogo Que Voce Respeita Memes23 maio 2024
Aquele Jogo Que Voce Respeita Memes23 maio 2024 -
 Pizza Siciliano (Sicilian Style Pizza) Tasty Kitchen: A Happy Recipe Community!23 maio 2024
Pizza Siciliano (Sicilian Style Pizza) Tasty Kitchen: A Happy Recipe Community!23 maio 2024 -
 Louveira goleia de novo e garante classificação para 2ª fase do Paulista feminino23 maio 2024
Louveira goleia de novo e garante classificação para 2ª fase do Paulista feminino23 maio 2024 -
 One punch man season 3 Anime one punch man, Saitama one punch man, Saitama one punch23 maio 2024
One punch man season 3 Anime one punch man, Saitama one punch man, Saitama one punch23 maio 2024 -
 Hogwarts™ Moment: Herbology Class 76384 | Harry Potter™ | Buy online at the Official LEGO® Shop US23 maio 2024
Hogwarts™ Moment: Herbology Class 76384 | Harry Potter™ | Buy online at the Official LEGO® Shop US23 maio 2024
